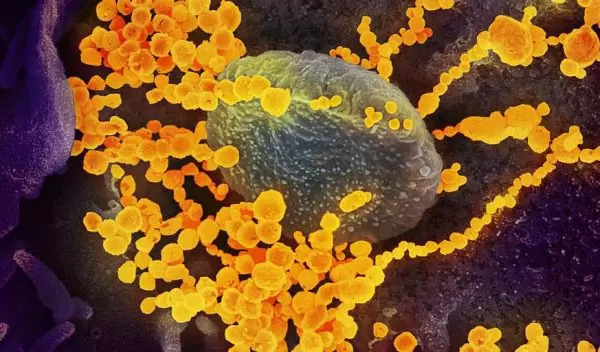
A machine-learning approach to finding treatment options for Covid-19
When the COVID-19 pandemic struck in early 2020, doctors and researchers rushed to find effective treatments. There was little time to spare. "Making new drugs takes forever," says Caroline Uhler, a computational biologist at the Massachusetts Institute of Technology. "Really, the only expedient option is to repurpose existing drugs."
Uhler's team has now developed an approach based on machine learning to identify drugs already on the market that could potentially be repurposed to fight COVID-19, particularly in the elderly. The system accounts for changes in gene expression in lung cells caused by both the disease and aging.
That combination could allow medical experts to more quickly seek drugs for clinical testing in elderly patients, who tend to experience more severe symptoms. The researchers pinpointed the protein RIPK1 as a promising target for COVID-19 drugs, and identified three approved drugs that act on the expression of RIPK1.
The findings of the U.S. National Science Foundation-funded research appear in the journal Nature Communications.
The researchers zeroed in on the most promising drug-repurposing candidates in three broad steps. First, they generated a large list of possible drugs by using a machine-learning technique called an autoencoder. Next, they mapped the network of genes and proteins involved in both aging and SARS-CoV-2 infection.
Finally, they used statistical algorithms to understand causality in that network, allowing them to pinpoint "upstream" genes that caused cascading effects throughout the network. In principle, drugs targeting those upstream genes and proteins should be promising candidates for clinical trials.
"This breakthrough is an example of how funding basic research over time gives us the knowledge we need to respond to the problems of the day," says Gabor Szekely, a program director in NSF's Division of Mathematical Sciences.
